

CACHE is short for Critical Assessment of Computational Hit-finding Experiments and is an initiative led by the Structural Genomics Consortium (SGC). The challenge focuses on providing unbiased experimental feedback on hit compounds and enhance research through an open-science approach.
.jpg)
We are honoured to be one of the 23 applicants accepted to the challenge; all participants accepted to the challenge are now published on the challenge website:
https://cache-challenge.org/challenges/finding-ligands-targeting-the-tkb-domain-of-cblb/computational-methods
For ANYO Labs this is an opportunity to highlight the AI-based drug discovery methods developed by Professor Leif A Eriksson and Dr S. Jalil Mahdizadeh to rapidly identify small molecular hit compounds for protein targets.
In this challenge the goal is to find ligands targeting the TKB domain of CBLB, an E3 ubiquitin protein ligase involved in several cancer types. The target is the subject of several patented compounds and a well understood problem; however, the crystal structure (PDB code 8GCY) reveals the inactive state of the protein. The challenge requires participants to identify potential binders for the same pocket as the co-crystalized ligand in the provided PDB file using their in-silico techniques. The identified hits should be structurally novel (compared to the co-crystalized ligand) with a Kd value below 30 uM.
The organization also provided a list of 895 chemicallyrelated compounds with confirmed inhibitory effect on the protein that act as abenchmark for newly identified hits.
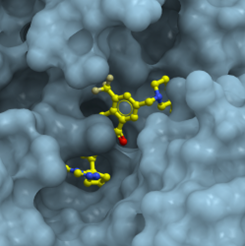
The new hits could encompass any chemical structure, and for the first “hit Identification” step of the challenge, the organizer recommended participants to make use of the Enamine REAL library (https://enamine.net/compound-collections/real-compounds/real-database) to increase the number of compounds being tested and reduce costs for custom synthesis.
In our application you can find how we utilise our proprietary ML-pipeline (i-TripleD) to indicate novel hits using generative AI. We aimed to deploy our full pipeline starting from only the PDB file to deeply evaluate the characteristics of the binding pocket, start generating compounds and filter out only the top scoring ones based on predicted Kd values. In this stage, target to hit, the interest is in predicting and filtering by binding affinity only. Since our methodology does not require conformational sampling, we expect our approach to be most computationally efficient while still retaining high accuracy. The initial plan was to generate de novo compounds and then perform a similarity search on the recommended Enamine library to find close analogues to be purchased cheaply in this initial stage of hit-screening.

To start a project, a protein visualization tool is used to understand the target and the co crystalized ligand. Literature is also reviewed to “re-score” some of the known binders and to validate the target in our models. Afterwards, screening setup is initiated. Because of the extensive timeline of the challenge, we in this case deployed i-TripleD on one computational node utilising one A-100 GPU for 168h (1week).
During this time, the code was able to generate and screen approximately one billion compounds, 785 of which were predicted to bind with sub-nanomolar affinity. However,when a similarity check was performed towards the Enamine REAL library, nothing was available. The same was true when checking the MolPort Library. After some research, it was evident that i-TripleD, which explores the complete binding site, had generated larger compounds than what was available in the Enamine library. This indicates that the pocket we are trying to target has more possible interactions than what is utilized by the co-crystalized ligand.
When brought up to the Cache challenge representatives, they did confirm that the binding pocket is indeed large and “might not be ideally compatible with a cut-off of 500 Da” but believed that there would still be potential hits to be found in the provided Enamine library (since the distribution for the CBLD inhibitors with IC50 <5 micromolar show that over half have a MW <500 Daltons).
Our belief is that the best hits would have been outside the Enamine library. In this case though ,the screening method was switched entirely towards the two trenches of the Enamine REAL library containing the largest compounds (~1 billion compounds HAC 29-38). This screening took 36 hours and only 58 276 hits were found with predicted pKd above 8 (binding affinity better than 10 nM). After additional filtering, we ended up choosing ~40 compounds out of the Enamine REAL library for further testing by the Organization as opposed to up to 100 that we could have selected.
Amidst the ongoing AI-driven transformation in the field of drug discovery, we find ourselves continually uncovering the possibilities and constraints of our methods and those of the pharmaceutical industry. In this particular scenario, trusting AI entirely would have necessitated the synthesis of custom compounds to identify the most promising hit compounds. However, we opted to focus solely on the molecules available in the Enamine library.
This decision underscores the importance of initiating true denovo chemistry efforts even at the early stages of development. A tenfold enhancement in binding affinity at this juncture can significantly influence whether a candidate progresses smoothly through subsequent development phases. Given more resources, it would be fascinating to assess how our best-predicted binders would perform in this assay. Fortunately, there is a second round of experimental testing scheduled for next year, offering the prospect of testing some of these compounds.
Nonetheless, we view the current project's trajectory a success, as it showcases the company's agility and its ability to adapt tools to evolving situations. If it proves successful despite the constraints we faced, this challenge also presents a unique opportunity to contribute to the scientific community and gain significant exposure at this stage.
We are currently awaiting the results from the experimental testing which will be published in February. We have the hope that there are several potent binders among the 40 candidates selected from our screen, which will highlight the unique capabilities of i-TripleD.